 
Often warning messages are caused by dependencies failing to install. 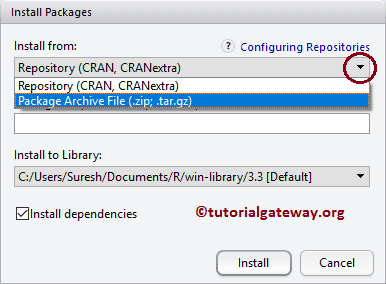
Setting R_REMOTES_NO_ERRORS_FROM_WARNINGS="false" will cause warning messages during calls to install.packages() to become errors. Setting R_REMOTES_STANDALONE="true" forces remotes to work in standalone mode and avoid loading its optional dependencies (curl, git2 and pkgbuild currently. 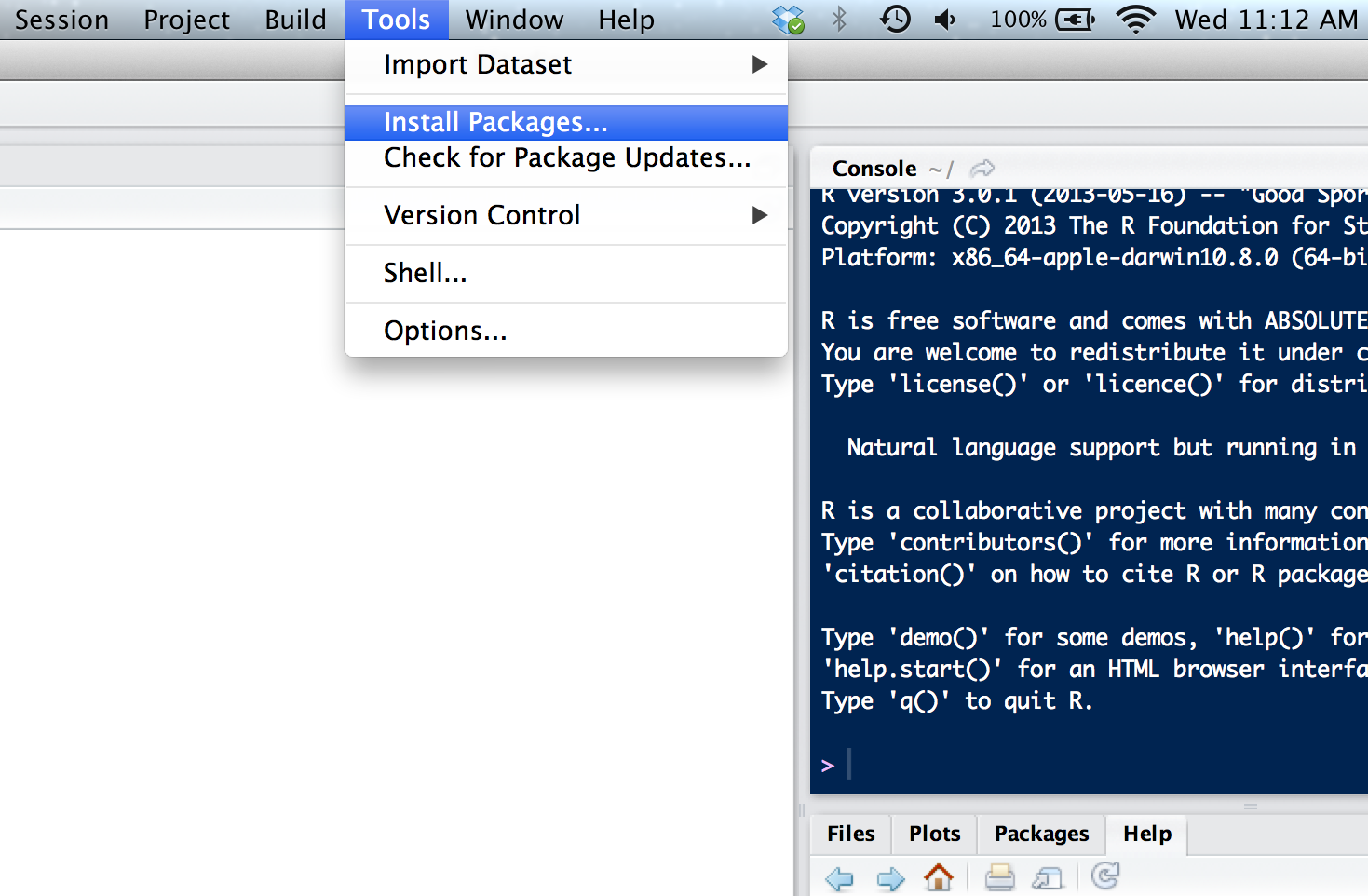
For example, you can set R_REMOTES_UPGRADE="always" to upgrade dependent packages without asking the user. The R_REMOTES_UPGRADE environment variable can be used to set a default preferred value for the upgrade = argument accepted by the various install_*() functions. The R_BIOC_VERSION environment variable can be used to force a Bioconductor version. (The BioC_mirror option takes precedence over this.) The most common way is to use the CRAN repository, then you just need the name of the package and use the command install.packages ('package'). So, for publicly available packages, this means to what repository it belongs. The R_BIOC_MIRROR environment variable can be used to specify an alternative Bioconductor mirror. Installing R Packages From CRAN How you can install an R package will depend on where it is located. The GITHUB_PAT environment variable is used as the default GitHub personal access token for all GitHub API queries. The BITBUCKET_USER and BITBUCKET_PASSWORD environment variables are used for the default Bitbucket user name and password, in install_bitbucket() 
0 Comments
|
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
